# Dell Hack AI Challenge
## Retrieval Augmented Generation system for Biolofical Species Taxonomy Classification
## Table of Contents
1. [Dependencies](#dependencies)
2. [Project Motivation](#motivation)
3. [Running the model in NVIDIA AI Workbench](#running)
4. [Features and Functionality](#features)
5. [Files Description](#description)
6. [Results](#results)
7. [Acknowledgements](#acknowledgements)
### Dependencies
All the following libraries are needed to implement this project:
**Python**
**Ollama**
**Llama-index**
**Bio**
**Pandas**
**Huggingface-hub**
**pyarrow**
It is also highly recommended to use NVIDIA AI Workbench to streamline the process of setting up the environment.
### Project Motivation
This project is an effort to demonstrate a proof of concept of the utility of RAG for academic research, leveraging the developmental aid of NVIDIA AI workbench. We have built a retrieval augmented generation system for the NCBI species taxonomy classification database making use of the Tree Index from Llamaindex.
### Running the model in NVIDIA AI Workbench
1. Install and configure AI Workbench locally and open up AI Workbench. Select a location of your choice.
2. Fork this repo into your own GitHub account.
3. Inside AI Workbench:
a. Click Clone Project and enter the repo URL of your newly-forked repo.
b. AI Workbench will automatically clone the repo and build out the project environment, which can take several minutes to complete.
c. Open the model in Jupyter notebook and experiment!
### Features and Functionality
- Hierarchical Structure: Biological classifications follow a nested hierarchy (Domain, Kingdom, Phylum, Class, Order, Family, Genus, Species). This naturally maps to a tree structure.
- Efficient Queries: Tree-based indexes allow for efficient querying of hierarchical data. For example, finding all species within a genus or all genera within a family becomes a simple tree traversal operation.
- Relationship Preservation: The parent-child relationships in the taxonomy are preserved in a tree structure, making it easy to navigate up and down the classification hierarchy.
- Range Queries: Tree-based indexes are particularly good for range queries, which could be useful for retrieving all taxa between certain classification levels.
- Prefix Matching: Many tree-based indexes support efficient prefix matching, which is useful for partial name searches in taxonomic data.
- Scalability: Tree-based indexes can handle large datasets efficiently, which is crucial given the vast number of species and taxonomic groups.
- Updates: While not frequently needed in established taxonomies, tree structures can accommodate updates (like adding new species or reclassifications) relatively easily.
- LCA (Lowest Common Ancestor) Queries: Tree structures make it efficient to find the lowest common ancestor of two species, which is a common operation in taxonomic research.
### Files Description
*final_notebook.ipynb* : Notebook with detaield explanation of all the steps involved in our project
*preBuild.bash* : Script containing preBuild instructions for installing Ollama
*species_list.txt* : Species for which we are extracting taxonomy and building the RAG application
### Results
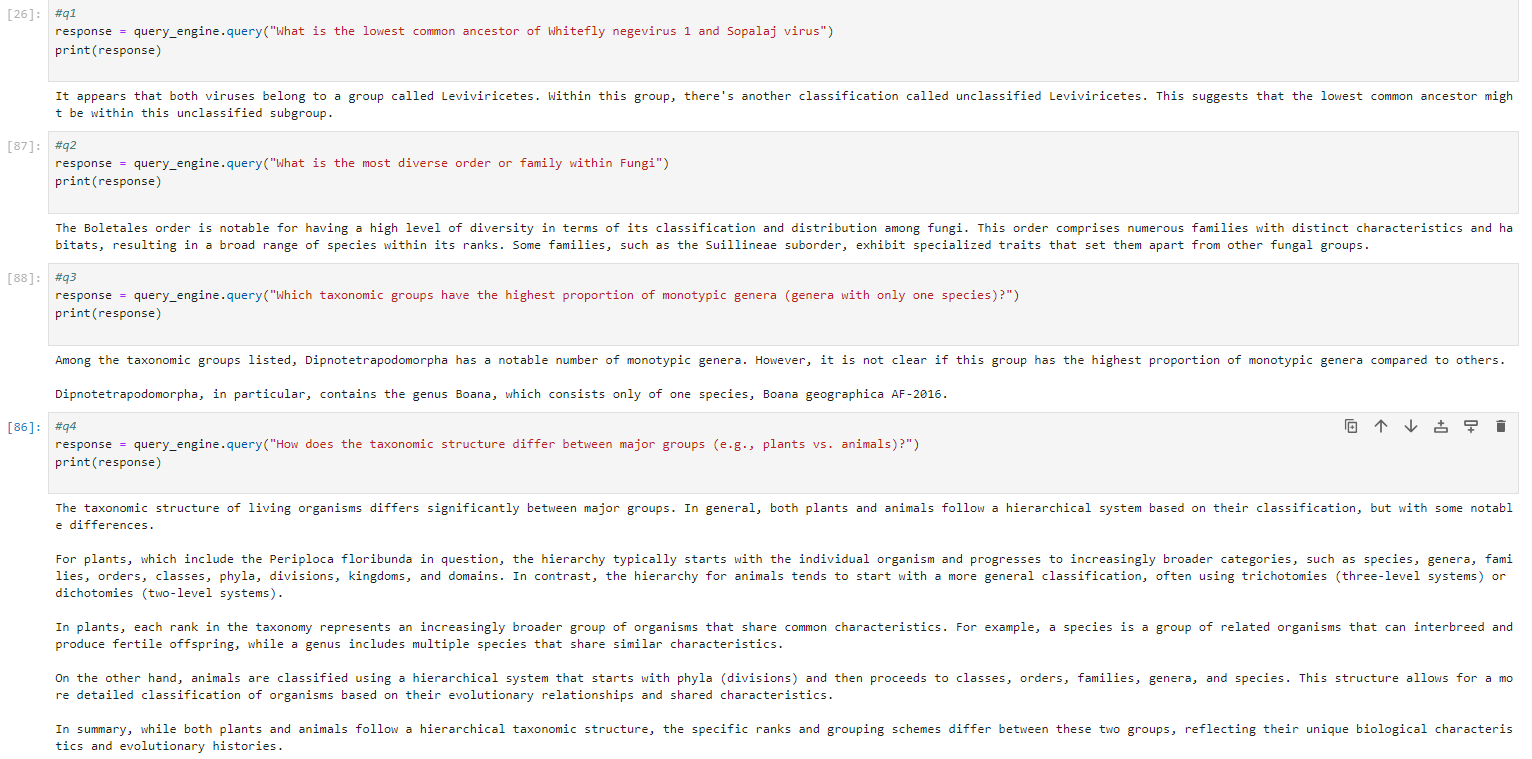
### Acknowledgements
We would like to thank NCBI for providing access to the information necessary for this project and hope it would be of great value for researchers in this domain.