OT_ICA_high_dim_bad_ic_test.ipynb
Optimal Transport in liner ICA
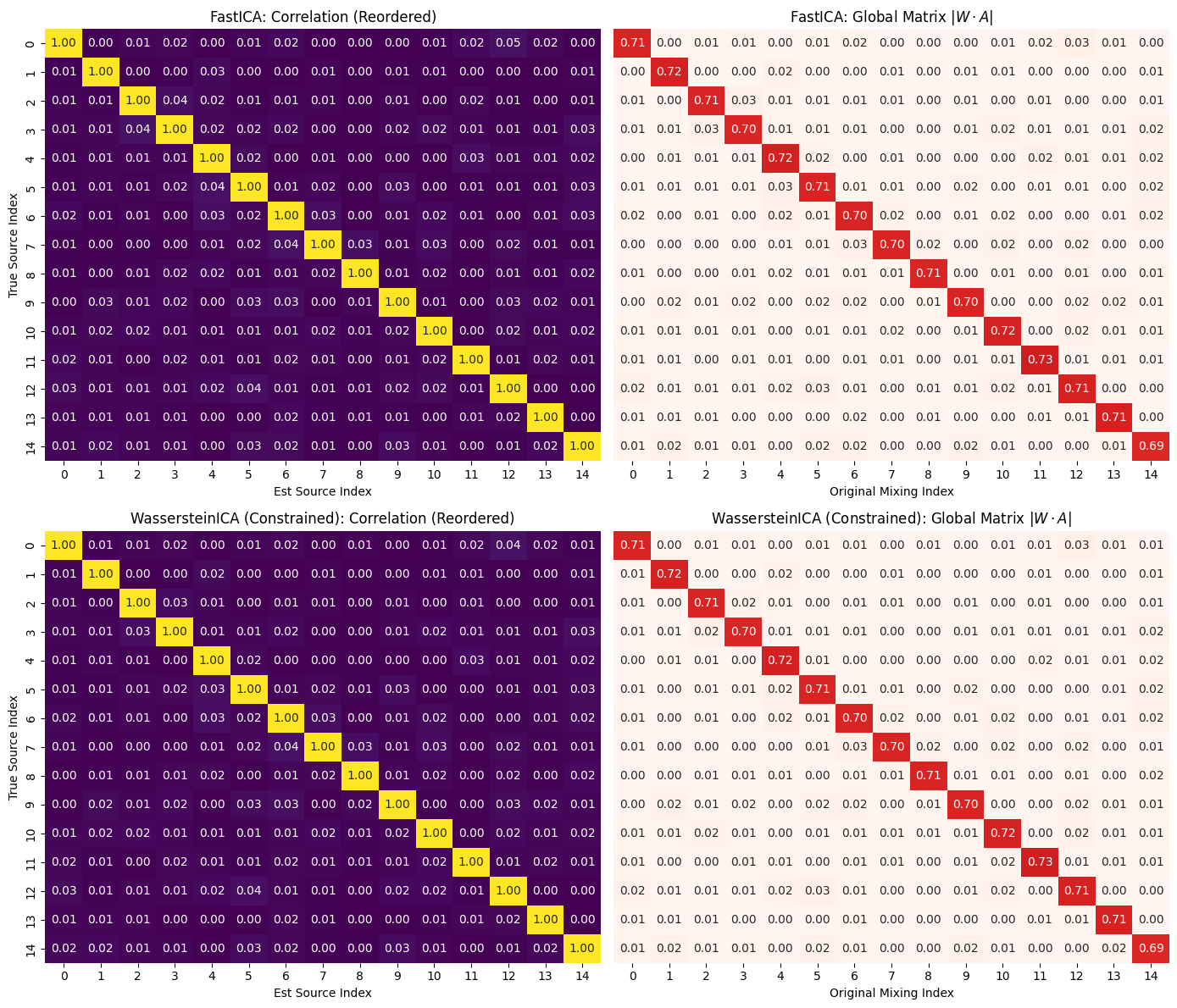
We find the ICs in high (10-15) dimensional case we plot the mixing unmixing matrix, compare the result to Fast ICA, and then transfer the ICs obtained in constrained space to unconstrained space to check that this is only iterative error accumulation and the OT algorithm is actually able to find areas where actual ICs reside.
import numpy as np
import torch
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
from scipy.optimize import linear_sum_assignment
from sklearn.decomposition import FastICA
from IPython.display import display
from wasserstein_ica import WassersteinICA
# ==========================================
# 1. Helper Functions
# ==========================================
def get_whitening_matrix(X_torch):
n_samples = X_torch.shape[1]
X_centered = X_torch - torch.mean(X_torch, dim=1, keepdim=True)
cov = torch.matmul(X_centered, X_centered.t()) / (n_samples - 1)
D, E = torch.linalg.eigh(cov)
D_inv_sqrt = torch.diag(1.0 / torch.sqrt(D + 1e-5))
W = torch.matmul(D_inv_sqrt, E.T)
return W.cpu().numpy()
def match_sources(S_true, S_est):
"""
Matches estimated sources to true sources using Hungarian algorithm.
Returns reordered S_est, the correlation matrix, and the permutation indices.
"""
n_dim = S_true.shape[0]
corr_mat = np.zeros((n_dim, n_dim))
for i in range(n_dim):
for j in range(n_dim):
corr_mat[i, j] = np.abs(np.corrcoef(S_true[i], S_est[j])[0, 1])
row_ind, col_ind = linear_sum_assignment(-corr_mat)
return S_est[col_ind], corr_mat[:, col_ind], col_ind, row_ind
def plot_ica_performance(corr_mat, global_mat, title_prefix, axes_row):
"""
Plots Correlation and Global Matrix for a specific method.
"""
# Correlation Matrix
sns.heatmap(corr_mat, ax=axes_row[0], cmap="viridis", vmin=0, vmax=1, annot=True, fmt=".2f", cbar=False)
axes_row[0].set_title(f"{title_prefix}: Correlation (Reordered)")
axes_row[0].set_ylabel("True Source Index")
axes_row[0].set_xlabel("Est Source Index")
# Global Matrix |WA|
sns.heatmap(global_mat, ax=axes_row[1], cmap="Reds", vmin=0, vmax=1, annot=True, fmt=".2f", cbar=False)
axes_row[1].set_title(f"{title_prefix}: Global Matrix $|W \\cdot A|$")
axes_row[1].set_yticks([])
axes_row[1].set_xlabel("Original Mixing Index")
def refine_subspace(ica, w_init, good_vectors, lr=0.1, max_iter=200):
"""
Optimizes w_init using Projected Gradient Descent.
Instead of a penalty, we explicitly project the gradient
to be orthogonal to the 'good_vectors' at every step.
"""
w = w_init.clone().detach().to(ica.X.device)
w.requires_grad_(True)
# Create the matrix of "fences" (Good Vectors)
if good_vectors is not None and len(good_vectors) > 0:
# Stack and normalize just in case
V = torch.stack(good_vectors).detach().to(ica.X.device)
V = V / torch.norm(V, dim=1, keepdim=True)
else:
V = None
# Use SGD for stability in this specific subspace check
for i in range(max_iter):
# 1. Compute Wasserstein Gradient
loss = -ica.wasserstein2_analytical(w)
loss.backward()
with torch.no_grad():
grad = w.grad
# 2. Subspace Projection (The "Hard" Fence)
# Remove any part of the gradient that points towards a Good Vector
# Formula: grad_new = grad - sum( (grad . v) * v )
if V is not None:
overlaps = torch.matmul(V, grad) # Shape (num_good,)
# Projection: sum over all good vectors
# We reshape overlaps to (num_good, 1) to broadcast over V (num_good, dim)
correction = torch.sum(overlaps.unsqueeze(1) * V, dim=0)
grad = grad - correction
# 3. Tangent Projection (Stay on the Sphere)
# Remove part of gradient parallel to w itself
grad = grad - torch.dot(grad, w) * w
# 4. Step
w += lr * grad
w /= torch.norm(w) # Renormalize
w.grad.zero_()
return w
# ==========================================
# Experiment : Error accumulation in High-Dim ICA
# ==========================================
def run_high_dim_experiment(n_dim=10, n_samples=2000):
print(f"--- Running {n_dim}D Experiment (N={n_samples}) ---")
# Generate Data
np.random.seed(42)
torch.manual_seed(42)
# Generate True Sources (Laplace)
sources = [np.random.laplace(0, 1, n_samples) for _ in range(n_dim)]
S_true = np.stack(sources)
# Mixing
A_true = np.random.randn(n_dim, n_dim)
X_np = A_true @ S_true
X_torch = torch.tensor(X_np, dtype=torch.float32)
# Reconstruct S_true_scaled for fair correlation check
# (Since A is random, S is not strictly perfectly recovered without scale/perm fix)
# We use the raw S_true for correlation which is scale invariant.
# 2. FastICA Run
print("Running FastICA...")
fast_ica = FastICA(n_components=n_dim, max_iter=2000, tol=1e-3, random_state=42)
S_fast = fast_ica.fit_transform(X_np.T).T
W_fast_total = fast_ica.components_ # This is unmixing matrix
# Match & Sort FastICA
S_fast_ordered, corr_fast, col_ind_fast, _ = match_sources(S_true, S_fast)
P_fast = np.abs(W_fast_total @ A_true)
P_fast = P_fast[col_ind_fast, :] # Reorder rows to look diagonal
# 3. WassersteinICA Run
print("Running WassersteinICA (Constrained)...")
ica = WassersteinICA(X_torch)
ica.whiten()
# Phase 1: Deflation (Standard SGD)
extracted_ws = []
for _ in range(n_dim):
prev = torch.stack(extracted_ws) if extracted_ws else None
# Note: calling without init_w as per current class design
w, _ = ica.optimize_wasserstein2(prev_components=prev, max_iter=200, n_restarts=50)
extracted_ws.append(w)
W_deflation = torch.stack(extracted_ws)
# Phase 2: Symmetric Refinement (L-BFGS, Standard W2)
# We purposefully turn OFF Sinkhorn as it is slow
W_sphere = ica.optimize_symmetric(
n_components=n_dim,
init_w=W_deflation,
max_iter=100,
lr=1.0,
optimizer='lbfgs',
use_sinkhorn=False
)
print("WassersteinICA complete")
# Reconstruct
W_white = get_whitening_matrix(X_torch)
W_wass_total = W_sphere.cpu().numpy() @ W_white
S_wass = W_wass_total @ X_np
# Match & Sort Wasserstein
S_wass_ordered, corr_wass, col_ind_wass, row_ind_wass = match_sources(S_true, S_wass)
P_wass = np.abs(W_wass_total @ A_true)
P_wass = P_wass[col_ind_wass, :]
# Visualization
fig, axes = plt.subplots(2, 2, figsize=(14, 12))
plot_ica_performance(corr_fast, P_fast, "FastICA", axes[0])
plot_ica_performance(corr_wass, P_wass, "WassersteinICA (Constrained)", axes[1])
plt.tight_layout()
plt.show()
# 5. The Subspace Check
print("\n--- Subspace Refinement Check ---")
print("Refining 'Bad' components while forcing orthogonality to 'Good' components (> 0.95)...")
diagonals = np.diag(corr_wass)
# Identify Good and Bad indices
good_indices = np.where(diagonals >= 0.95)[0]
bad_indices = np.where(diagonals < 0.9)[0] # Raised threshold slightly to catch more
if len(bad_indices) == 0:
print("No components < 0.95 found. Perfect recovery!")
else:
# 1. Collect the Good Vectors (The "Fences")
good_vectors = []
for idx in good_indices:
original_idx = col_ind_wass[idx] # Map to W_sphere row
good_vectors.append(W_sphere[original_idx])
results_table = []
for i in bad_indices:
original_idx = col_ind_wass[i]
current_corr = diagonals[i]
w_bad = W_sphere[original_idx]
# 2. Refine with Subspace Constraint
w_refined = refine_subspace(ica, w_bad, good_vectors, lr=0.1)
# Check 1: Did refining the bad vector work?
s_new_est = (w_refined.cpu().detach().numpy() @ W_white) @ X_np
new_corr = np.abs(np.corrcoef(S_true[i], s_new_est)[0, 1])
# --- OPTIONAL: The "Random Kick" ---
# If the bad vector was truly stuck, try a fresh random search in the subspace
if new_corr < 0.95:
w_random = torch.randn_like(w_bad)
w_random /= torch.norm(w_random)
w_refined_rand = refine_subspace(ica, w_random, good_vectors, lr=0.1)
s_rand_est = (w_refined_rand.cpu().detach().numpy() @ W_white) @ X_np
rand_corr = np.abs(np.corrcoef(S_true[i], s_rand_est)[0, 1])
# If random search worked better, keep that score
if rand_corr > new_corr:
new_corr = rand_corr
# Mark that we needed a restart
print(f" > Source {i}: Direct refinement failed ({new_corr:.2f}), but Random Restart found it ({rand_corr:.2f})!")
results_table.append({
"Source Idx": i,
"Orig Corr": current_corr,
"New Corr": new_corr,
"Recovered?": "YES" if new_corr > 0.95 else "NO"
})
# Display Results
df_check = pd.DataFrame(results_table)
display(df_check)
# Calculate stats
recovered_count = df_check[df_check["New Corr"] > 0.95].shape[0]
total_bad = len(bad_indices) # <--- DEFINE THIS VARIABLE
print(f"\nSummary: {recovered_count}/{total_bad} bad components were recovered in subspace refinement.")
if recovered_count == total_bad:
print("CONCLUSION: The Wasserstein objective is correct. The error was purely due to interference from other components (The Tower Problem).")
elif recovered_count > 0:
print("CONCLUSION: Partial Recovery. Orthogonality constraints were a major factor, but some landscapes may be too difficult.")
else:
print("CONCLUSION: No recovery. The issue likely lies in the loss landscape itself (local optima) rather than constraints.")
return ica, S_true, A_true, W_sphere, col_ind_wass
# 1. Run Experiment and Capture Objects
ica, S_true, A_true, W_sphere, col_ind_wass = run_high_dim_experiment(n_dim=15, n_samples=6000)
--- Running 15D Experiment (N=6000) ---
Running FastICA...
Running WassersteinICA (Constrained)...
WassersteinICA complete
--- Subspace Refinement Check ---
Refining 'Bad' components while forcing orthogonality to 'Good' components (> 0.95)...
No components < 0.95 found. Perfect recovery!
def check_oracle_stability(ica, S_true, A_true, W_alg, col_ind_alg):
"""
Initializes vectors exactly at the True Inverse (A_inv) and runs optimization.
Compares the W2 scores of the Oracle vs the Algorithm's results.
"""
print("\n--- Oracle Stability & Score Check ---")
# 1. Oracle Initialization: W_true = A_inv
A_inv = np.linalg.inv(A_true)
W_white_np = ica.W_white.cpu().numpy()
# Project A_inv onto the Whitened Sphere
W_white_inv = np.linalg.pinv(W_white_np)
W_oracle_sphere_np = A_inv @ W_white_inv
# Normalize rows to be unit norm (on the sphere)
norms = np.linalg.norm(W_oracle_sphere_np, axis=1, keepdims=True)
W_oracle_sphere_np = W_oracle_sphere_np / norms
W_oracle = torch.tensor(W_oracle_sphere_np, dtype=torch.float32).to(ica.X.device)
results = []
for i in range(W_oracle.shape[0]):
# Oracle Vector
w_oracle = W_oracle[i]
# Matching Algorithm Vector
w_alg_matched = W_alg[col_ind_alg[i]]
# Verify initial correlation of Oracle (Should be ~1.0)
s_init = (w_oracle.cpu().numpy() @ W_white_np) @ (A_true @ S_true)
init_corr = np.abs(np.corrcoef(S_true[i], s_init)[0, 1])
# --- CALCULATE W2 SCORES (Lower is Better) ---
# Note: We use the analytical method here
w2_oracle = ica.wasserstein2_analytical(w_oracle).item()
w2_alg = ica.wasserstein2_analytical(w_alg_matched).item()
# Run Unconstrained Refinement from the perfect spot
w_final = refine_subspace(ica, w_oracle, good_vectors=None, lr=0.01, max_iter=100)
s_final = (w_final.cpu().detach().numpy() @ W_white_np) @ (A_true @ S_true)
final_corr = np.abs(np.corrcoef(S_true[i], s_final)[0, 1])
results.append({
"Source": i,
"Start Corr": init_corr,
"End Corr": final_corr,
"Stable?": "YES" if final_corr >= init_corr - 0.01 else "DRIFT",
"Oracle W2": np.round(w2_oracle, 6),
"Algorithm W2": np.round(w2_alg, 6),
"W2 Difference": np.round(w2_alg - w2_oracle, 6) # Positive means Oracle is better (lower dist)
})
df = pd.DataFrame(results)
display(df)
# Summary of W2 gap
avg_diff = df['W2 Difference'].mean()
print(f"\nAverage W2 Score Gap: {avg_diff:.6f}")
if avg_diff > 0:
print("CONCLUSION: The Oracle solutions have strictly lower W2 distances (better scores).")
print("This proves the algorithm gets stuck in sub-optimal local minima during the search.")
# 2. Run the check
check_oracle_stability(ica, S_true, A_true, W_sphere, col_ind_wass)
--- Oracle Stability & Score Check ---
| Source | Start Corr | End Corr | Stable? | Oracle W2 | Algorithm W2 | W2 Difference | |
|---|---|---|---|---|---|---|---|
| 0 | 0 | 1.0 | 0.999841 | YES | 0.035298 | 0.035366 | 0.000068 |
| 1 | 1 | 1.0 | 0.999899 | YES | 0.039824 | 0.039766 | -0.000058 |
| 2 | 2 | 1.0 | 0.999633 | YES | 0.035829 | 0.036338 | 0.000509 |
| 3 | 3 | 1.0 | 0.999757 | YES | 0.038522 | 0.038651 | 0.000129 |
| 4 | 4 | 1.0 | 0.999816 | YES | 0.041695 | 0.041667 | -0.000028 |
| 5 | 5 | 1.0 | 0.999673 | YES | 0.038725 | 0.039105 | 0.000380 |
| 6 | 6 | 1.0 | 0.999629 | YES | 0.034730 | 0.035175 | 0.000446 |
| 7 | 7 | 1.0 | 0.999764 | YES | 0.037949 | 0.038264 | 0.000314 |
| 8 | 8 | 1.0 | 0.999864 | YES | 0.042369 | 0.042318 | -0.000051 |
| 9 | 9 | 1.0 | 0.999595 | YES | 0.042218 | 0.042876 | 0.000659 |
| 10 | 10 | 1.0 | 0.999866 | YES | 0.033049 | 0.033170 | 0.000120 |
| 11 | 11 | 1.0 | 0.999729 | YES | 0.035715 | 0.036074 | 0.000359 |
| 12 | 12 | 1.0 | 0.999589 | YES | 0.039492 | 0.040161 | 0.000669 |
| 13 | 13 | 1.0 | 0.999820 | YES | 0.036040 | 0.036113 | 0.000073 |
| 14 | 14 | 1.0 | 0.999671 | YES | 0.035988 | 0.036630 | 0.000641 |
Average W2 Score Gap: 0.000282
CONCLUSION: The Oracle solutions have strictly lower W2 distances (better scores).
This proves the algorithm gets stuck in sub-optimal local minima during the search.