OT_natural_grad_vs_FastICA.ipynb
Optimal Transport in linear ICA
In this notebook we test the performance of a OT Natural Gradient ICA versus Fast ICA over dimensions.
import numpy as np
import torch
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
from sklearn.decomposition import FastICA
from tqdm.notebook import tqdm
# Import your package
from wasserstein_ica import WassersteinICA
# ==========================================
# 1. Helpers & Data Generation
# ==========================================
def amari_error(W_est, A_true):
if W_est is None or np.any(np.isnan(W_est)):
return np.nan
P = np.abs(W_est @ A_true)
n = P.shape[0]
row_sum = np.sum(P, axis=1)
row_max = np.max(P, axis=1)
term1 = np.sum((row_sum / row_max) - 1)
col_sum = np.sum(P, axis=0)
col_max = np.max(P, axis=0)
term2 = np.sum((col_sum / col_max) - 1)
return (term1 + term2) / (2 * n)
def generate_dataset(n_dim, n_samples, seed=None):
if seed is not None:
np.random.seed(seed)
torch.manual_seed(seed)
# Standard Laplace (Super-Gaussian)
sources = [np.random.laplace(0, 1, n_samples) for _ in range(n_dim)]
S = np.stack(sources)
# Well-conditioned mixing matrix
cond_num = 1000
while cond_num > 100:
A = np.random.randn(n_dim, n_dim)
cond_num = np.linalg.cond(A)
X = A @ S
return torch.tensor(X, dtype=torch.float32), A
# ==========================================
# 2. Experiment Setup
# ==========================================
# Test a range of dimensions up to 30
DIMENSIONS = [n for n in range(5, 51, 5)] # Varying D
N_SAMPLES = 5000
N_TRIALS = 5 # Run multiple trials for error bars
print(f"--- FastICA vs. OT Fixed-Point Showdown ---")
print(f"Dimensions: {DIMENSIONS}")
print(f"Samples: {N_SAMPLES}")
print(f"Trials per dim: {N_TRIALS}")
results = []
--- FastICA vs. OT Fixed-Point Showdown ---
Dimensions: [5, 10, 15, 20, 25, 30, 35, 40, 45, 50]
Samples: 5000
Trials per dim: 5
# ==========================================
# 3. Main Loop
# ==========================================
for dim in tqdm(DIMENSIONS, desc="Dimensions"):
for trial in range(N_TRIALS):
# 1. Generate Data
X_torch, A_true = generate_dataset(n_dim=dim, n_samples=N_SAMPLES, seed=trial)
X_np = X_torch.numpy()
# 2. FastICA (Baseline)
try:
fast_ica = FastICA(n_components=dim, max_iter=2000, tol=1e-4, random_state=trial)
fast_ica.fit(X_np.T)
W_fast_total = fast_ica.components_
score_fast = amari_error(W_fast_total, A_true)
except Exception as e:
score_fast = np.nan
W_fast_total = None
results.append({'Dimension': dim, 'Method': 'FastICA', 'Amari Error': score_fast})
# 3. Initialize WassersteinICA
ica = WassersteinICA(X_torch)
ica.whiten()
W_white_np = ica.W_white.cpu().numpy()
# 4. Wasserstein Fixed-Point (Cold Start - Random)
try:
# We don't pass init_w, so it starts random
W_sphere_cold = ica.optimize_fixed_point(n_components=dim, max_iter=100, tol=1e-5)
W_wass_cold_total = W_sphere_cold.cpu().numpy() @ W_white_np
score_cold = amari_error(W_wass_cold_total, A_true)
except Exception as e:
score_cold = np.nan
results.append({'Dimension': dim, 'Method': 'W-ICA (Cold Start)', 'Amari Error': score_cold})
# 5. Standalone WassersteinICA (Phase 1 + Natural Gradient Polish)
try:
# PHASE 1: Robust Deflation (Find the mountains)
extracted_ws = []
for _ in range(dim):
prev = torch.stack(extracted_ws) if extracted_ws else None
# Using 50 restarts to guarantee we avoid local minima
w, _ = ica.optimize_wasserstein2(prev_components=prev, max_iter=200, n_restarts=50)
extracted_ws.append(w)
W_deflation_init = torch.stack(extracted_ws)
# PHASE 2: Natural Gradient Polish over GL(n)
# We feed the Phase 1 output into optimize_symmetric using the 'natural' optimizer
W_natural_unmixed = ica.optimize_symmetric(
n_components=dim,
max_iter=200,
lr=0.5, # Slightly lower learning rate recommended for stability in GL(n)
init_w=W_deflation_init,
optimizer='natural',
penalty_weight=1.0 # Controls the log|det W| volume-preserving penalty
)
# CRITICAL ADDITION: Because Natural Gradient optimizes over the non-orthogonal
# General Linear Group GL(n), we must symmetrically orthogonalize it ONE FINAL TIME
# before calculating the Amari error against the whitened data space.
W_final_orthogonal = ica._symmetric_decorrelation(W_natural_unmixed)
W_wass_total = W_final_orthogonal.cpu().numpy() @ W_white_np
score_wass = amari_error(W_wass_total, A_true)
except Exception as e:
print(f"Failed at dim {dim}: {e}")
score_wass = np.nan
results.append({'Dimension': dim, 'Method': 'W-ICA (Natural Gradient)', 'Amari Error': score_wass})
Dimensions: 0%| | 0/10 [00:00<?, ?it/s]
df = pd.DataFrame(results)
plt.figure(figsize=(10, 6))
# seaborn's lineplot automatically plots the mean and a confidence interval for the trials
sns.lineplot(data=df, x='Dimension', y='Amari Error', hue='Method', marker='o', linewidth=2.5)
plt.title("Amari Error vs. Dimension\n(FastICA vs. Wasserstein Fixed-Point)", fontsize=14)
plt.ylabel("Amari Error (Lower is Better)", fontsize=12)
plt.xlabel("Number of Dimensions", fontsize=12)
plt.grid(True, linestyle='--', alpha=0.7)
#plt.yscale('log') # Log scale is often better for Amari Error to see small polish improvements
plt.tight_layout()
plt.show()
# Display mean scores table
display(df.groupby(['Dimension', 'Method'])['Amari Error'].mean().unstack().round(4))
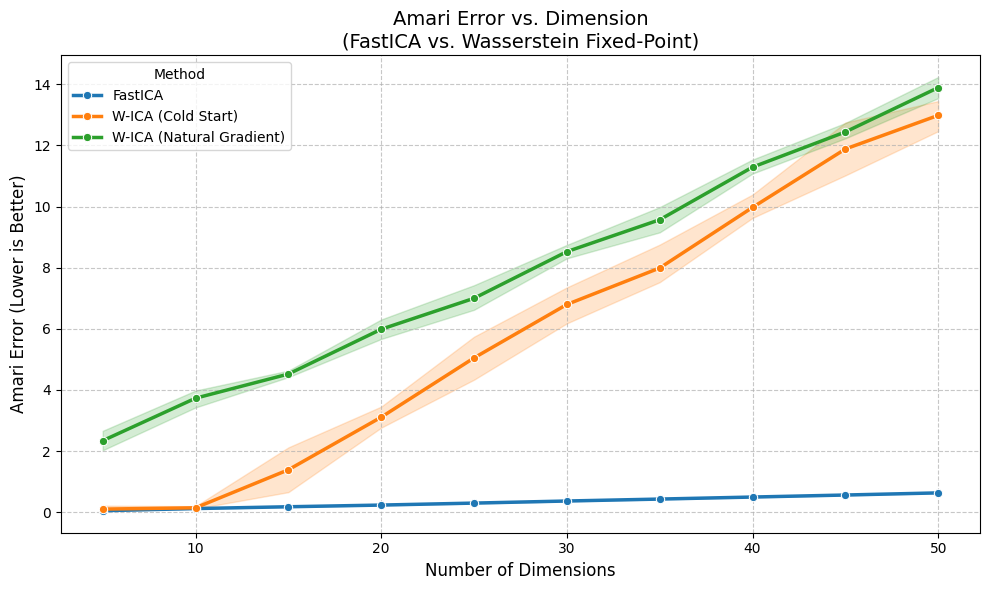
| Method | FastICA | W-ICA (Cold Start) | W-ICA (Natural Gradient) |
|---|---|---|---|
| Dimension | |||
| 5 | 0.0482 | 0.1003 | 2.3385 |
| 10 | 0.1183 | 0.1428 | 3.7281 |
| 15 | 0.1771 | 1.3867 | 4.5186 |
| 20 | 0.2324 | 3.1077 | 5.9845 |
| 25 | 0.2971 | 5.0471 | 6.9972 |
| 30 | 0.3653 | 6.7954 | 8.5238 |
| 35 | 0.4287 | 7.9887 | 9.5671 |
| 40 | 0.4952 | 9.9702 | 11.2877 |
| 45 | 0.5612 | 11.8817 | 12.4396 |
| 50 | 0.6310 | 12.9816 | 13.8901 |